|
George Shaikovski I am a PhD student at the University of Cambridge, supervised by Mateja Jamnik and supported by Gates Cambridge scholarship. I work on visuospatial reasoning, planning, and reinforcement learning. Traditional RL excels in learning implicit spatial reasoning (think AlphaZero or DQN beating Atari), however it doesn't generalise beyond its input distribution and baked-in goals. Language models generalise reasoning to novel scenarios, arbitrary goals, and unseen environments via universality of language. However they struggle to perform visuospatial reasoning beyond simple problems. Bridging the gap between the two is the primary goal of my research. Before Cambridge, I was a research engineer at Paige for 5 years. We built large vision encoders to learn efficient microscopic tissue representations for computational pathology, and vision-language generative models for specimen-level representations and diagnostic report generation. Check out our models on Huggingface. Email / Google Scholar / Twitter / LinkedIn / Github |

|
Research |
|
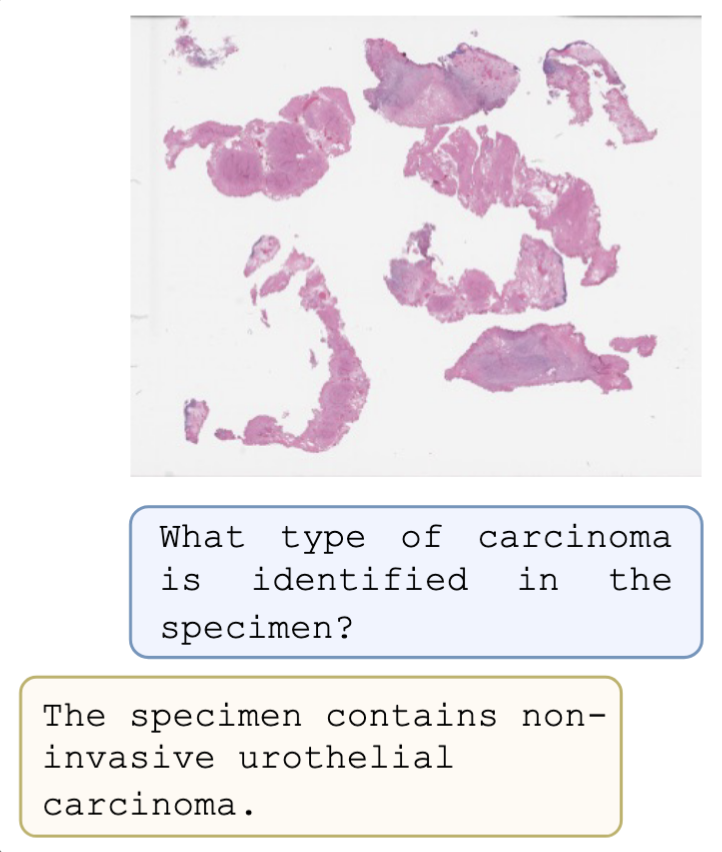
PRISM2: Unlocking Multi-Modal General Pathology AI with Clinical Dialogue
Eugene Vorontsov, George Shaikovski, Adam Casson, Julian Viret, Eric Zimmermann, Neil Tenenholtz, Yi Kan Wang, Jan H. Bernhard, Ran A. Godrich, Juan A. Retamero, Jinru Shia, Mithat Gonen, Martin R. Weiser, David S. Klimstra, Razik Yousfi, Nicolo Fusi, Thomas J. Fuchs, Kristen Severson, Siqi Liu arXiv, 2025 arXiv Utility of vision-language pre-training for histopathology had been limited by small datasets and insufficient grounding in diagnostic reasoning. We scaled PRISM to 700k specimen-report pairs and expanded supervision signal to clinical-dialogue, aligning histomorphologic features with diagnostic language. The method matched and even exceeded cancer-detection performance of clinical-grade products without additional training. |
|
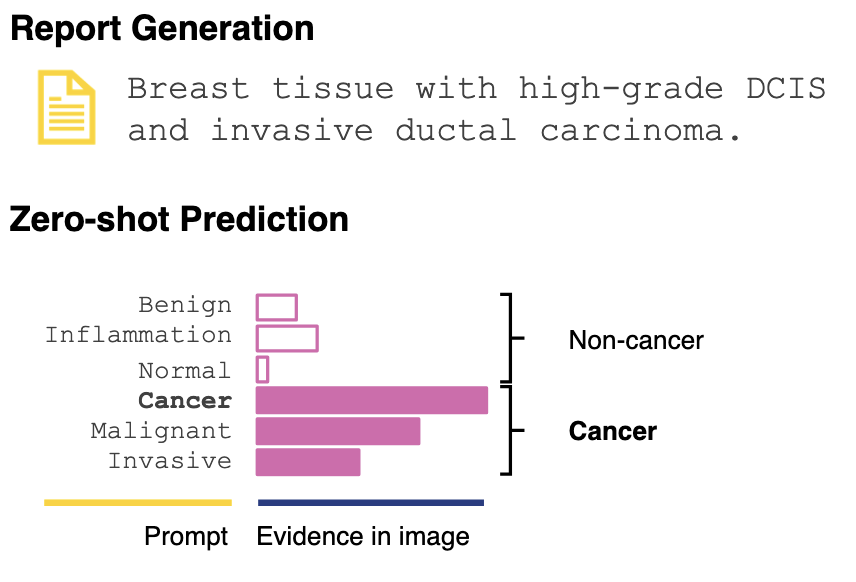
PRISM: A Multi-Modal Generative Foundation Model for Slide-Level Histopathology
George Shaikovski, Adam Casson, Kristen Severson, Eric Zimmermann, Yi Kan Wang, Jeremy D. Kunz, Juan A. Retamero, Gerard Oakley, David Klimstra, Christopher Kanan, Matthew Hanna, Michal Zelechowski, Julian Viret, Neil Tenenholtz, James Hall, Nicolo Fusi, Razik Yousfi, Peter Hamilton, William A. Moye, Eugene Vorontsov, Siqi Liu, Thomas J. Fuchs arXiv, 2024 arXiv / model Clinical pathology reports are at the specimen level, yet foundation models are limited to small patches within specimens. We developed PRISM, a vision-language model which aggregates representations across 100k patches to match detailed clinical diagnosis extracted from the reports, thus creating a denser supervision signal. The method enables 1) zero-shot cancer detection via text prompts and 2) data-efficient transfer of specimen-level representations to biomarker prediction, which outperforms supervised baselines with as little as 10% of the training data. |
|
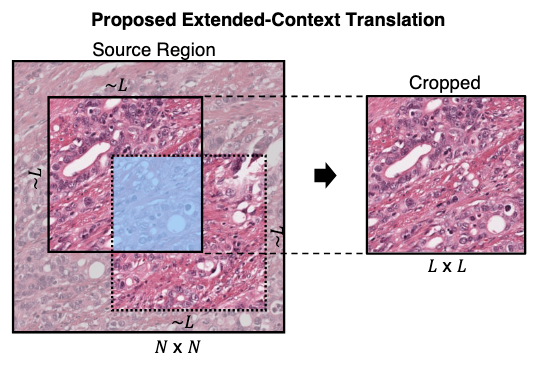
Virchow2: Scaling Self-Supervised Mixed Magnification Models in Pathology
Eric Zimmermann, Eugene Vorontsov, Julian Viret, Adam Casson, Michal Zelechowski, George Shaikovski, Neil Tenenholtz, James Hall, David Klimstra, Razik Yousfi, Thomas Fuchs, Nicolo Fusi, Siqi Liu, Kristen Severson arXiv, 2024 arXiv / model We investigate the most effective way to scale foundation models for histopathology. Prior work scaled along individual axes of data and model size, however an exploration of domain-specific changes to the algorithm was lacking. We introduced new data augmentations and a regularisation objective tuned to pathology data, and demonstrate their advantages over the vanilla DINOv2 method in a large ablation study. We then successfully scaled Virchow to 3.1M whole slide images and 1.9B parameters, achieving state-of-the-art on 12 tile-level tasks. |
|
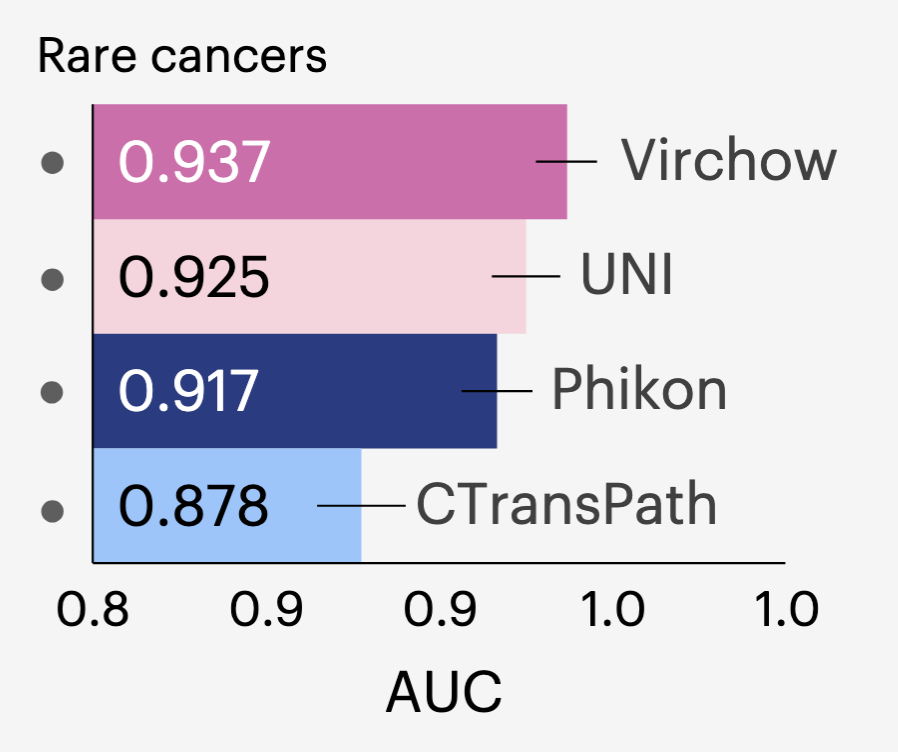
A Foundation Model for Clinical-Grade Computational Pathology and Rare Cancers Detection
Eugene Vorontsov, Alican Bozkurt, Adam Casson, George Shaikovski, Michal Zelechowski, Kristen Severson, Eric Zimmermann, James Hall, Neil Tenenholtz, Nicolo Fusi, Ellen Yang, Philippe Mathieu, Alexander van Eck, Donghun Lee, Julian Viret, Eric Robert, Yi Kan Wang, Jeremy D. Kunz, Matthew C. H. Lee, Jan H. Bernhard, Ran A. Godrich, Gerard Oakley, Ewan Millar, Matthew Hanna, Hannah Wen, Juan A. Retamero, William A. Moye, Razik Yousfi, Christopher Kanan, David S. Klimstra, Brandon Rothrock, Siqi Liu, Thomas J. Fuchs Nature Medicine, 2024 paper / model Computational pathology models must generalise across diverse tissue types and rare cancers, which requires large-scale diverse datasets. However, prior foundation models had not been scaled to the data volumes needed for clinical-grade performance on rare tissue types. We trained Virchow, a ViT-H DINOv2 model, on 1.5M whole slide images, the first histopathology foundation model at this scale, enabling clinical-grade tile-level feature extraction with strong generalisation to rare tissues and cancers. |
MiscellaneousI enjoy sailing small boats. My dream is to get an International Yachtmaster license. |
|
Design from Jon Barron's website. |